The main feature of the Library of Identification Resources is the description of the identification key (or matrix, reference, etc.). This description should on its basis specify when the key can or should be used. I have initially split this description into the taxonomic coverage and the ‘scope’. The latter includes life stage and sex but also some restrictions on the taxonomic coverage that are more difficult to characterize, like “species that cause galls on plants in the Rosa genus” or “species that live in aquatic environments”.
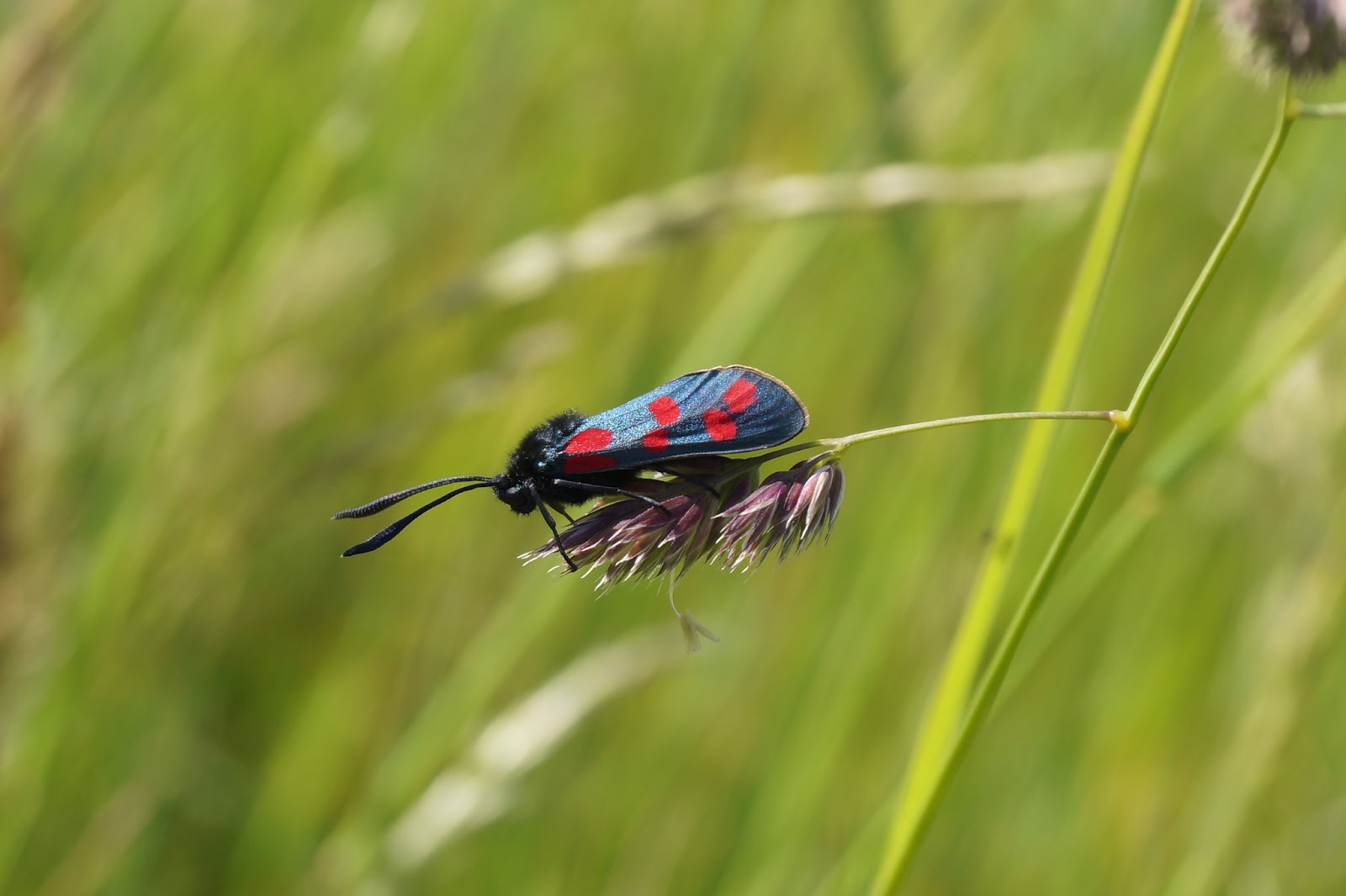

Zygaena filipendulae (Linnaeus, 1758)
Simple solution
For the taxonomic coverage sensu stricto, I started with the parent taxon, e.g. the family or genus. However, many keys do not treat all the species in a family, and are instead limited by a geographic scope. This geographic scope should clearly also be included. Then there are more casual keys that can be very useful but may not be complete even at the time of publication, either excluding some rare species or only including some common species. This can be detailed (to some extent) with an incomplete/complete switch. Finally, although many keys are for species, there are some keys primarily for identifying genera, families, or other ranks. Below are some example works where these aspects do or do not apply.
| Title |
Parent taxon |
Geographic scope |
Incomplete |
Target rank |
| B460: A revision of the world Embolemidae (Hymenoptera Chrysidoidea) |
Embolemidae |
— |
— |
— |
| B1: Identification Key to the European Species of the Bee Genus Nomada Scopoli, 1770 (Hymenoptera: Apidae), Including 23 New Species |
Nomada Scopoli, 1770 |
Europe |
— |
— |
| B81: Key to some European species of Xylomyidae |
Xylomyidae |
Europe |
Yes |
— |
| B63: MOSCHweb - Interactive key to the genera of the Palaearctic Tachinidae (Insecta: Diptera) |
Tachinidae |
Palaearctic realm |
— |
genus |
If only it were that simple
It became clear quite quickly that this is not enough. For one, parent taxon, geographic scope, and target rank should be able to contain multiple values. Additionally, as we saw before, some taxonomic coverages like “gallers on Rosa sp.” cannot be captured with these parameters unless the “Parent taxon” list gets very long and detailed.
Another, more common problem is that even the combination of a parent taxon and a geographic scope is hardly specific enough to be able to say whether an identification key is reliable and complete for a situation. Species are discovered, migrate, emigrate, and become extinct. Taxa are moved to different genera, families, and orders. In B659: Orthoptêres et Dermaptêres (Faune de France 3) from 1922, the order Orthoptera also contains Dictyoptera, currently a superorder consisting of cockroaches, stick insects, praying mantises, and more. This is a big problem too. Key questions that should be answerable are “How well does a British key apply to a Dutch observation?” and “How well does a British key from 1950 apply to a British observation from 2020?”
To account for the changes within higher-level taxa you might want to make a list of species that are included in the keys. For B81 for example, that could look like this:

https://purl.org/identification-resources/resource/B81:1
This gives a very clear image of the taxonomic coverage of the key: it includes three species, Salva marginata, S. varia and Xylomya maculata. The inclusion of the species in this key has a permalink, and the taxon is linked to a GBIF identifier. The list of species can then be matched (especially with the identifiers) to current checklists for the region that the observation was made in. More on that later.
This solution can easily be used for more complex taxonomic coverages, like the aforementioned key for species that cause galls on Rosa species. The ‘species’ list can also be a taxon list and have e.g. genera as the lowest rank. Another advantage is that there is no longer a need to explicitly specify the geographic scope, whether or not the key was known to be incomplete at the time of writing, and what the target taxon rank of the key is. In addition, this specific implementation also allows for multiple keys per work, which has various uses.
Two purposes
One problem that still comes up is that these taxon lists have two purposes:
- They describe which taxa can be distinguished with the key.
- They describe which taxa are considered when making the key.
These two are at odds. In a key to genera it would be simple to make a list of these genera, fulfilling (1) but not (2): if a species in of those genera gets moved to a different one, or if an additional species in one of the existing genera appears, the key ‘breaks’. The latter could be fixed by just listing all species but at that point you might as well fix both problems with (2) by grouping the species by genus. To simultaneously fulfill purpose (1) and (2) it is necessary to divide the taxonomic tree into units that can be distinguished by the key.
Even when fulfilling purpose (1) it is still possible to partially accomplish (2) in keys for e.g. families that key out to species. If, when making the species list, higher ranks such as genera are included, the key to genera can be validated according to (2) without having to go into more detail than the key. Therefore it is important to capture the entire taxonomic tree as presented in the work.
Another thing that might be useful to record is the synonyms. Many of the more academic works publish along with the key a series of descriptions including synonyms. If the status of that synonymic taxon is now different from the status in the key, this may be of importance to the results of the identification. It comes down to the same split of purposes: (1) no distinction is made in the key between the two taxa but (2) both taxa are, in theory, considered. Either way it is important to list both in some way.
If only it were that simple
At this point it became clear which taxa are necessary to include in such a list. But how to describe the taxa in those lists? This is an (enormous) rabbit hole in and of itself, and if I had more experience I could make a Falsehoods Programmers Believe About Names-style post about it. It starts with the normal parts: every taxon has a name, an author, a year, and a rank. That rank can be kingdom, phylum, class, order, family, genus, or species. But there is a seemingly endless list of increasingly obscure ranks, sometimes only in use by one or a few authors (what is a “stirps”??); taxa can have multiple authors and different ways of presenting this (et/and/& and et al. or listing everyone); the names themselves can be spelled in different ways and include or omit initials; the year can be very unclear, especially for taxa published in older works published over multiple years; and the scientific names themselves can have different spellings, capitalizations, and hyphenations as well.
As with personal names, the ‘trick’ is to trust the source to some extent, and avoid focussing on the specifics of the scientific name. After all, there is no reason to make the computer understand the scientific names. The goal is to allow for matching (and there are ways to do that without trying to standardize everything) and, secondarily, to present the names in such a way that humans can understand them (and the original key already did that).
Measuring applicability (more problems)
Now, with a complete and structured taxonomic tree, the question becomes how to check whether it applies to a certain observation. The idea is to compare the taxon list to a list of all the species that a certain observation could be. But where does such a list come from? The simple solution would be a checklist: a professional, thorough list of all the species that are known to occur in a certain area.
If such a checklist is not (easily) available (I am not aware of a global database of checklists, let alone in a standardised format), it might be tempting to create one from a map of observations from e.g. GBIF. The advantage of this is that you have, to some extent, global coverage and a standardised format. However, this has some caveats. The correctness of the resulting checklist is entirely dependent on the quality and quantity of the observations in the region. To get a higher sensitivity it may be useful to instead make a checklist of a larger, encompassing region, but this lowers the specificity. A species occurring in Belgium might only have recorded observations in the Netherlands on GBIF. The same goes for the time scale. There might be museum specimens of species that are now extinct in the region, but there might also be rare species that are only seen every 50 years or so.
A good addition (or a risky alternative) to a checklist would be a measure of the (relative) abundance of species in the region, and weighing the taxon lists of the keys according to that. This prioritises keys that include more common species over keys that include more rare species. Of course, the key of common species can be wrong about an observation of a rare species. Another problem is how to determine the (relative) abundance. Again it is tempting to derive this from GBIF observations, but a species with a lot of observations is not necessarily more common. It might be the focus of an observation campaign, or national attention of a different kind, it might sit still more often, it might be easier to recognize, or it might even be easier to identify to a species level.
An additional possibility of measuring applicability is bringing the scope restrictions mentioned at the start into this. Apart from belonging to a certain taxon, the observed organism also has a biological sex, a life stage, and more characteristics that may restrict which keys can be used. To compare these characteristics between keys and organisms they need to be described with a common, consistent vocabulary. iNaturalist has such a vocabulary (the “Annotations”), but there might be more suitable ones out there somewhere.
(For this it might also be useful to map keys that distinguish castes of ants, males/females of solitary bees, or life stages of shield bugs. How do you model that? Not in the same way as described above, that is for sure.)
Using the data
As I teased in the previous blog post about the Library of Identification Resources, I have started to collect this data, and attempted to recruit some others to contribute as well. All works which have their keys (and matrices, checklists, descriptions) mapped out can be found by searching for “TRUE” in the “Tax. data extr.” field. To get from the work page to a taxa list, look for the row titled “Resources” in the first table on the page. If available it lists the individual resources in the work for which the taxonomic data is extracted.
The pages for the taxonomic data of the individual resources contains some basic information about the work (as well as a link), as well as the page numbers of the resource within the work if available. There is also some info on the resource, derived from the same info in the work unless different for that resource. Then there is a list of taxa, displayed as a tree. Each taxon has a rank, an anchor, and if available a link to the corresponding taxon in GBIF.
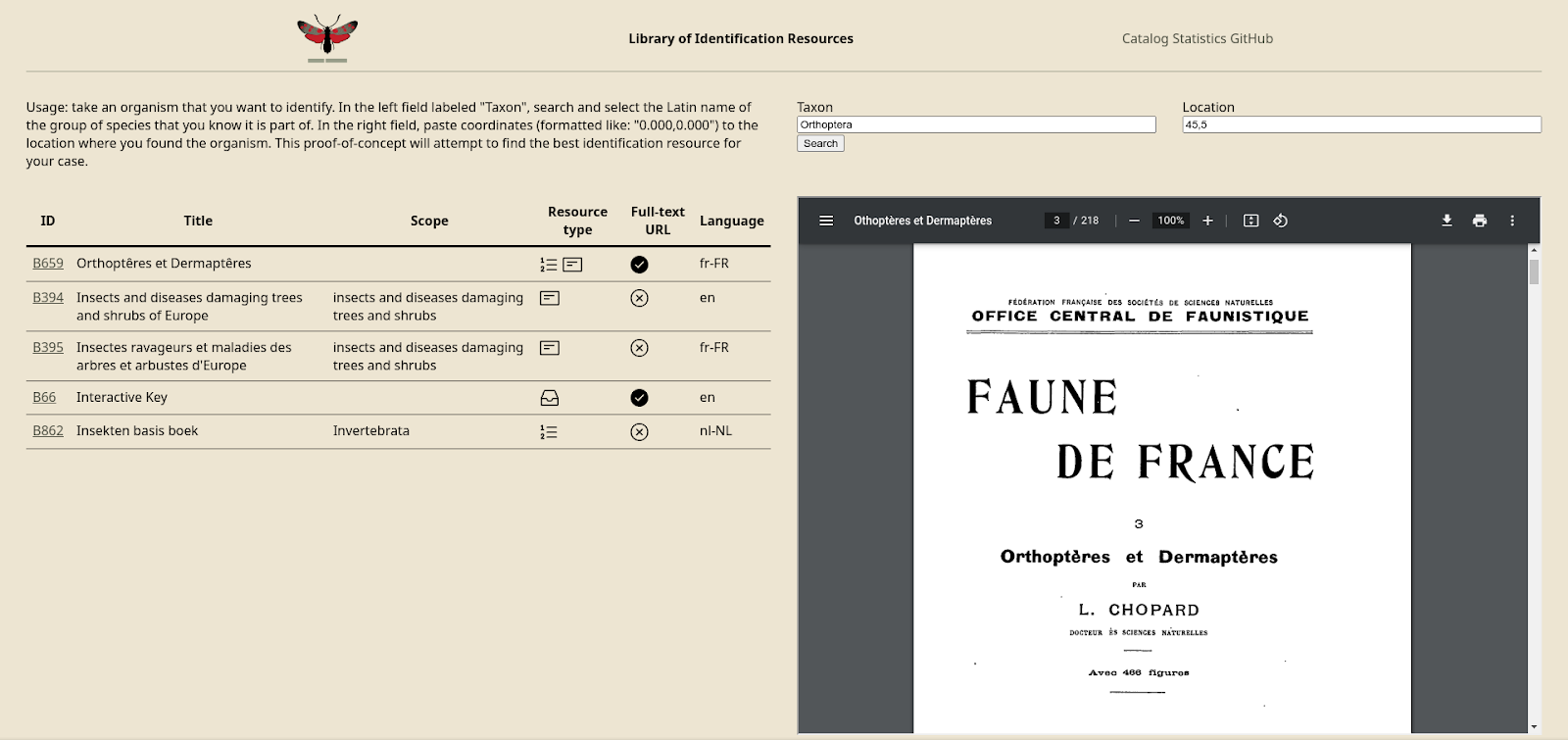
The data is also used in the proof-of-concept app that I introduced in the previous blog post. Searching for a taxon and coordinates will now query GBIF for observations in that taxon in the country encompassing the coordinates, and match this with the GBIF identifiers in the taxon lists. It displays the relative amount of taxa in the ‘checklist’ that are also in the key as a percentage and a small pie chart. It does not yet deal with synonyms.














 Ginkgo biloba leaves (source:
Ginkgo biloba leaves (source: